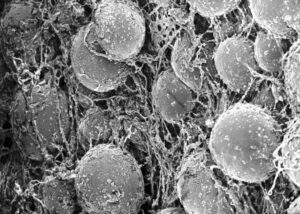
Why a liver on a chip?
The liver is part of the digestive tract, and the main organ involved in the metabolism of xenobiotics, thus in toxicological studies (either fundamental or during the drug development path), it is mandatory to test the chemical on the liver. Currently in use models are badly predictive of what is then observed in humans. Liver toxicity is one of the major causes of arrest during clinical phases of the development of a new drug, with the so-called Drug-Induced Liver Injury (DILI).
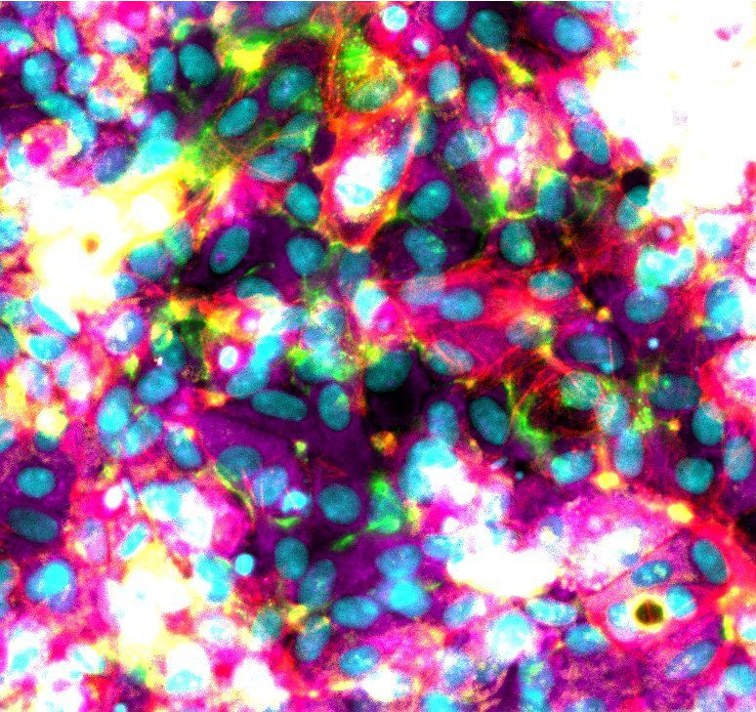
How to culture vascularized & immunocompetent 3D models in a standard Multiwell
What has already been achieved in the organ on a chip liver field?
The first model that might be closed to a liver on a chip is an assembly of micro-pillars in microwells (532 wells plate) with a 60nL capacity (1). Hepatic cells are suspended in a Matrigel and then placed on the micro-pillar. This model is compatible with high-throughput screening methods (HTS).
Regarding HTS methods, it is now possible to use micropatterning techniques (soft lithography) with co-cultured cell types in a 384 wells plate (Bhatia et al). The main issue remains the use of PDMS which absorbs some drugs.
Jumping to liver on a chip, a team in Pittsburg University (Lansing & Vernetti) has developed a liver containing hepatocytes, endothelial cells, Kupfer cells, and stellate cells (a) (2). Cells are coming from a biopsy or of nonusable organ donation. This system is used for hepatotoxicity assessment (dead and bursting of cells) with the aim to better predict and prevent the occurrence of DILI. Emulate has developed a liver-on-chip containing several cell types. This chip is currently used by the FDA (b). The Healy lab has developed a liver on chip derived from iPS cells with 4 weeks viability (c).
BioMEMS has developed a microfluidic device aiming at mimicking hepatic sinusoids aiming at better prevent DILI. This device contains the same cell types as Lansing’s one (3).
A collaborative project involving Columbia University, MIT, Boston University, and the Wyss Institut combined iPS differentiated in a 3D scaffold mimicking the environment of a hepatic lobule. This device is then coupled to a similar device containing heart cells or blood microvessels. The platform might be used to model diseases and during drug screening steps.
A team hosted at Harvard University (Khademhosseini et al) has bio-printed a hepatic spheroid on a microfluidic device (4).
Columbia University and MIT are collaborating on a hepatic spheroid on a chip device as well.
Globally, 2 types of platforms are used: statics platform (without perfusion system) and platform with a perfusion system. The last is used by Griffith et al (MIT), Toh, and colleagues (they generate a concentration gradient of a given chemical). Hµrel company also developed a platform with a perfusion system on a mono-layer of hepatic cells to evaluate the elimination of a given pharmaceutical, then extended to be compatible with co-culture.
What’s next in the liver-on-a-chip field?
The main issue with the “liver” organ on a chip is the bile ducts that have not been recreated yet. Once done, connected liver-on-a-chip with gut on a chip will give a more global vision about the way a chemical is washed out the body.
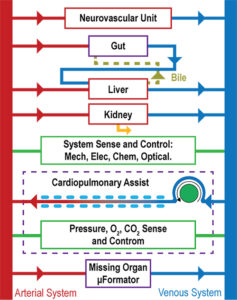
To go further simple liver-on-a-chip, scientists are developing PBPK (physiology-based pharmacokinetic) approaches to model what could be the whole organism reaction.
References
- Trends Biotechnol. 2017 Feb;35(2):172-183. doi: 10.1016/j.tibtech.2016.08.001. Epub 2016 Sep 2. Engineered Liver Platforms for Different Phases of Drug Development. Ware BR, Khetani SR
- Exp Biol Med (Maywood). 2016 Jan; 241(1): 101–114. A human liver microphysiology platform for investigating physiology, drug safety, and disease models. Lawrence A Vernetti, Nina Senutovitch, Robert Boltz, Richard DeBiasio, Tong Ying Shun, Albert Gough, and D Lansing Taylor
- http://www.biomemsrc.org/research/cell-tissue-microengineering/liver-on-a-chip
- Biofabrication. 2016 Jan 12;8(1):014101. doi: 10.1088/1758-5090/8/1/014101. A liver-on-a-chip platform with bioprinted hepatic spheroids. Bhise NS1, Manoharan V, Massa S, Tamayol A, Ghaderi M, Miscuglio M, Lang Q, Shrike Zhang Y, Shin SR, Calzone G, Annabi N, Shupe TD, Bishop CE, Atala A, Dokmeci MR, Khademhosseini
And also :
Miniature liver on a chip could boost US food safety
Meet Chip: Liver (from NIH)
A perfusion incubator liver chip for 3D cell culture with application on chronic hepatotoxicity testing
FAQ
The liver is the main organ involved in the metabolism of xenobiotics, which are foreign substances. It is therefore mandatory to test new chemicals on the liver during toxicological studies and drug development. A significant problem is that the models currently in use are badly predictive of the responses that are then observed in humans. Liver toxicity, known as Drug-Induced Liver Injury (DILI), is one of the major causes for the arrest of a new drug’s development during the clinical phases. New models are needed to better assess hepatotoxicity and to more accurately predict and prevent the occurrence of DILI.
The first model described as being close to a "liver on a chip" was an assembly of micro-pillars within microwells. This was a 532-well plate where each well had a 60nL capacity. In this system, hepatic cells were suspended in a Matrigel and subsequently placed onto the micro-pillars. This model is compatible with high-throughput screening (HTS) methods. It is also possible to use micropatterning techniques like soft lithography to create co-cultures of different cell types in a 384-well plate format. A main issue with some of these systems, however, remains the use of PDMS, which is a material that absorbs some drugs.
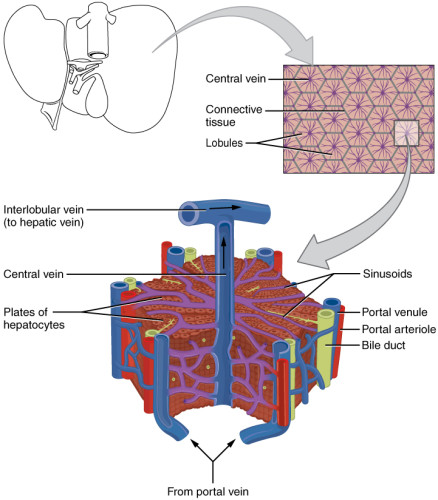
More advanced liver models have been developed to include multiple cell types. One such liver model, from Pittsburg University, contains hepatocytes, endothelial cells, Kupfer cells, and stellate cells. The cells for this system are sourced from biopsies or non-usable organ donations, and the model is used for hepatotoxicity assessment. Another microfluidic device from BioMEMS aims to mimic hepatic sinusoids and contains the same cell types. Other approaches use iPS cells that are differentiated in a 3D scaffold, mimicking the environment of a hepatic lobule. This device can be coupled to other devices containing heart cells. A hepatic spheroid has also been bio-printed onto a microfluidic device.
Globally, two types of platforms are used for these models. The first type is a static platform. These systems are used without a perfusion system. The second type is a platform that includes a perfusion system. This latter type is used by researchers at MIT, who have developed a system with a perfusion system on a mono-layer of hepatic cells. This setup is used to evaluate the elimination of a pharmaceutical. This platform was later extended to be compatible with co-culture. Other researchers have also used perfusion systems. For example, Toh and colleagues have used one to generate a concentration gradient of a specific chemical.
A main issue with the "liver" organ-on-a-chip models that have been developed is related to a missing component. The bile ducts, which are part of the liver’s digestive function, have not been recreated yet in these systems. Once this is successfully done, the next step would be to connect the "liver on a chip" with a "gut on a chip". Such a connection would provide a more global vision. This would help researchers understand the way a chemical is processed and washed out of the body. To advance beyond simple models, scientists are also developing physiology-based pharmacokinetic (PBPK) approaches to model the reaction of the whole organism.