For the first time Zhuo Du et al. correlate mRNA and protein expression in C. elegans embryo to build the first embryo protein mapping.
After the generated single-cell mRNA expression atlas of Jonathan S. Packer (2019) in C. elegans embryo, a mapping of proteins expression to better understand biological processes, especially during embryogenesis of C. elegans, has been made by Zhuo Du et al.. A representation encompassing 266 transcription factors allows them to establish a regulatory hierarchy between many lineages, tissues and transcription factors in spatiotemporal fate patterning; also highlighting new functions of certain transcription factors. Since the protein level cannot be directly correlated to the expression of an mRNA.
Check out, how they have tackled this challenge.
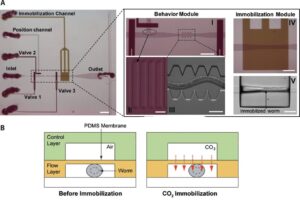
Ultra fast temperature shift device for in vitro experiments under microscopy
Abstract
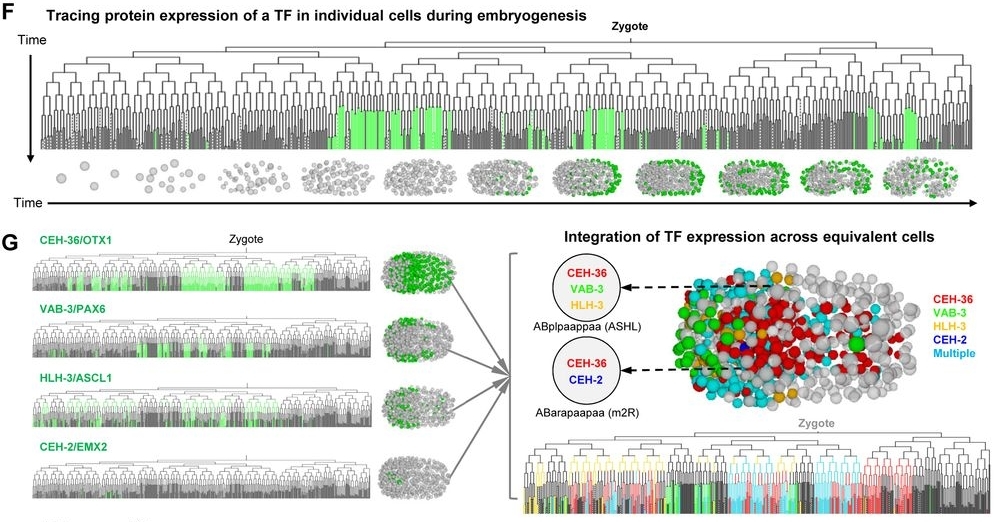
A high-resolution protein atlas is essential for understanding the molecular basis of biological processes. Using protein-fusion reporters and imaging-based single-cell analyses, we present a protein expression atlas of C. elegans embryogenesis encompassing 266 transcription factors (TFs) in nearly all (90%) lineage-resolved cells. Single-cell analysis reveals a combinatorial code and cascade that elucidate the regulatory hierarchy between a large number of lineage-, tissue-, and time-specific TFs in spatiotemporal fate patterning. Guided by expression, we identify essential functions of CEH-43/DLX, a lineage-specific TF, and ELT-1/GATA3, a well-known skin fate specifier, in neuronal specification; and M03D4.4 as a pan-muscle TF in converging muscle differentiation in the body wall and pharynx. Finally, systems-level analysis of TF regulatory state uncovers lineage- and time-specific kinetics of fate progression and widespread detours of the trajectories of cell differentiation. Collectively, our work reveals a single-cell molecular atlas and general principles underlying the spatiotemporal patterning of a metazoan embryo.
References
- Zhuo DU et al., biorxiv.org (2020)
FAQ
A new protein atlas has been presented. It details protein expression during the embryogenesis of C. elegans. This high-resolution atlas is considered necessary for understanding the molecular basis of biological processes. It was created using protein-fusion reporters. Imaging-based single-cell analyses were also used. The atlas encompasses 266 different transcription factors (TFs). Data is included for nearly all (90 percent) of the lineage-resolved cells in the embryo. This work follows a previously generated single-cell mRNA expression atlas. This new protein map provides a single-cell molecular atlas. It also helps explain the general principles for the spatiotemporal patterning of a metazoan embryo.
A mapping of protein expression was made to better understand biological processes. This is especially true for those processes occurring during the embryogenesis of C. elegans. A single-cell mRNA expression atlas had been previously generated. However, the protein level in a cell cannot be directly correlated to the expression of an mRNA. Therefore, this new protein mapping was needed. A high-resolution protein atlas is considered necessary for understanding the molecular basis of these processes. The new atlas allows for a better comprehension of spatiotemporal fate patterning. This work helps to build a protein map for the embryo.
The single-cell analysis reveals a combinatorial code. A cascade is also shown. These findings explain the regulatory hierarchy that exists between many transcription factors (TFs). This hierarchy is observed between different lineages and tissues. It relates to how cell fate is patterned over time and space. A representation of 266 TFs allows this regulatory hierarchy to be established. The analysis of the TF regulatory state at a systems-level also uncovers other details. The kinetics of fate progression, which are specific to lineage and time, are shown. Widespread detours in the trajectories of cell differentiation were also found.
The atlas was used to guide further investigation. This led to the identification of new functions for certain transcription factors. Necessary functions were identified for CEH-43/DLX. This is a lineage-specific TF. Functions were also found for ELT-1/GATA3. ELT-1/GATA3 is a known skin fate specifier. Both of these transcription factors were found to have necessary functions in neuronal specification. The atlas also highlighted another transcription factor, M03D4.4. This was identified as a pan-muscle TF. It is involved in converging muscle differentiation. This differentiation occurs in both the body wall and the pharynx.