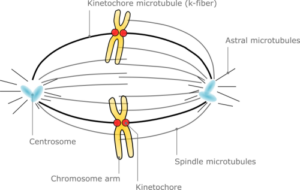
A fundamental breakthrough in the understanding of development through the realization of the complete transcriptome of C. elegans, from the embryo to the terminal state cell differentiation. Opening the gate for vertebrate development potential disease understanding.
During the development, expression of genes influences cell fate with impacts on the organism later in life. To better understand the fundamental mechanisms of development, gaining new knowledge about gene expression timing and pathways is necessary. Jonathan S. Packer et al. have identified almost all cells terminal and pre-terminal cell types by following the trajectories and temporal dynamics of their respective gene expression. This tracking during C. elegans embryogenesis has allowed these researchers to identify most single-cell transcriptomes. A breakthrough that they hope will lead to new projects and discoveries in vertebrate and potentially human.
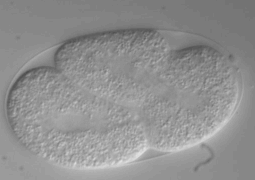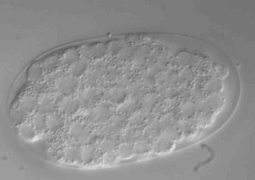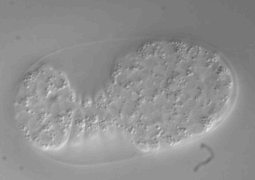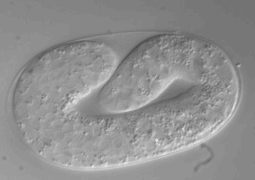
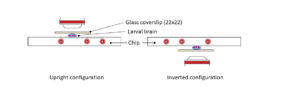
Ultra fast temperature shift device for in vitro experiments under microscopy
Abstract
Caenorhabditis elegans is an animal with few cells but a wide diversity of cell types. In this study, we characterize the molecular basis for their specification by profiling the transcriptomes of 86,024 single embryonic cells. We identify 502 terminal and preterminal cell types, mapping most single-cell transcriptomes to their exact position in C. elegans’ invariant lineage. Using these annotations, we find that (i) the correlation between a cell’s lineage and its transcriptome increases from middle to late gastrulation, then falls substantially as cells in the nervous system and pharynx adopt their terminal fates; (ii) multilineage priming contributes to the differentiation of sister cells at dozens of lineage branches; and (iii) most distinct lineages that produce the same anatomical cell type converge to a homogenous transcriptomic state.
References
FAQ
A study was conducted to characterize the molecular basis for cell specification in the C. elegans embryo. This animal is noted for having few cells but a wide diversity of cell types. The work involved profiling the transcriptomes of 86,024 single cells from embryos. This effort produced a molecular atlas. Researchers were able to identify 502 terminal and pre-terminal cell types. Most of the single-cell transcriptomes were successfully mapped to their exact position within the organism’s invariant lineage. This tracking provides new knowledge about gene expression timing. This information is required to better understand the mechanisms of development.
This research represents an important advance in the understanding of development. During an organism’s development, the expression of genes influences cell fate. This has later effects on the organism. Gaining new knowledge about the timing and pathways of gene expression is necessary to better understand these mechanisms. This study followed the trajectories and temporal dynamics of gene expression. This tracking allowed researchers to identify most single-cell transcriptomes and almost all terminal and pre-terminal cell types. It is hoped that this work will lead to new projects and discoveries. This may include a better understanding of vertebrate and human development or disease.
The annotations from the atlas were used for analysis. One finding relates to the correlation between a cell’s lineage and its transcriptome. This correlation was observed to increase. This increase occurs during the period from middle to late gastrulation. Following this period, the correlation falls substantially. This reduction happens as cells that are part of the nervous system and pharynx adopt their terminal fates. This demonstrates a dynamic relationship between cellular ancestry and gene expression. The mapping of single-cell transcriptomes to the organism’s known lineage was necessary for this observation.
The study’s annotations were used to examine how cell types are formed. It was found that most distinct lineages that produce the same anatomical cell type behave in a specific way. These different lineages were observed to converge. They arrive at a homogenous transcriptomic state. This finding means that even if cells originate from different ancestral paths, they become similar in their gene expression. This occurs as they form the same type of anatomical cell. Another finding was that multilineage priming contributes to the differentiation of sister cells. This happens at dozens of lineage branches. This work helps explain how a wide diversity of cell types can arise from a single embryo.