Introduction
Non-adherent cells are difficult to image because of movement. This is the case of living cells which are motile, but also of fixed cells which will move due to Brownian motion. While doing fluorescent microscopy, it is critical to prevent movement of cells to be imaged. This is particularly important when doing multichannel fluorescent microscopy, when there is a need to merge the two channels for colour overlays. There are several methods to prevent cell movement during cell imaging, and this Scientific Note focuses on agar pads.

Ultra fast temperature shift device for in vitro experiments under microscopy
What is an agar pad?
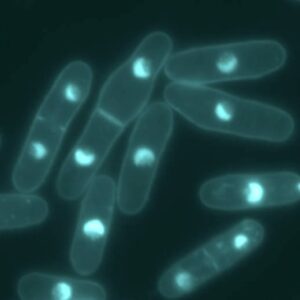
Agar pads prevent both motility of live cells and Brownian motion of fixed cells. This makes them the method of choice for live imaging of multiple model organisms: bacteria, yeast, worm embryos and adult worms, plant, zebrafish embryos. They can be coupled to additional methods to further enhance immobilization: for instance, the use of anesthetics for C. elegans.
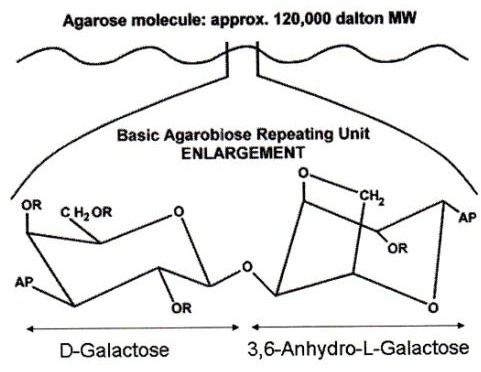
What is an agar pad? Although they are commonly referred as such, they are commonly made of agarose. Agar is a polysaccharide obtained from red algae, while agarose is the main component of agar, obtained after agar purification. Therefore, they are thin flat layers of an agarose matrix (see image). As a polymer, agarose creates a porous matrix where cells (or molecules for molecular biology applications) are trapped. However, nutrient and gas exchange occurs through the matrix making this method compatible with live imaging. No toxicity for the cells due to growth in agar has been described.
Agarose percentage to make pads can vary. Very standard percentages are from 0.5% to 2%. Higher percentages of agarose (up to 10%) have been used to immobilize adult worms, with the advantage of the lack of toxicity compared to other methods. Adult worms in agarose pads can be imaged for hours if the pad is properly sealed (avoiding desiccation) [3]. Replacement of agarose by other polysaccharides is as well possible. For instance, while 2% agar is the standard pad used for C. elegans embryo imaging, this can be replaced for 4% sucrose 0.1M NaCl for delicate early embryos or embryos sensitive to osmotic stress (Hyman Lab protocol, [2])
Melting and gelling temperatures depend on the type of agarose. Standard agarose is liquid after dissolution in water at high temperatures and solidifies around 35-45°C (depending on the concentration). Low melting agaroses remain fluid at much low temperatures (30°C).
What is an agar pad?
Agarose pads must be prepared in a way to ensure they are thin and flat, so they can be placed in a sandwich between a glass slide and a coverslip. Thickness of the pad will depend on the organism to be imaged. The following protocol shows how to prepare an 250uM thick pad by using simple and non-expensive laboratory materials: glass slides and laboratory tape. Note that thickness of the tape is a critical factor. If the tape is too thin, this will lead to very thin agar pads which will be very fragile and difficult to manipulate. (see our Agarose pad method for C. elegans temperature-sensitive embryos).
This mounting protocol is compatible as well with thicker agar pads. This could be needed if thicker specimens need to be imaged, like zebrafish embryos. The following video shows how an 1mm thick pad (percentage of agar 2%) can be thermalized using a CherryTemp device. The thickness of the agar slows down thermal transfer through the pad, although a homogeneous temperature across the pad is achieved after 30 seconds.The main limitations of agar pads for live cell imaging are the impossibility to change media (for instance to apply drugs), the risk of pad drying, and the limiting nutrient availability for long term imaging experiments. Microfluidic technologies, which ensure a more controlled environment for the cells, aim to overcome these limitations [4].
The main limitations of agar pads for live cell imaging are the impossibility to change media (for instance to apply drugs), the risk of pad drying, and the limiting nutrient availability for long term imaging experiments. Microfluidic technologies, which ensure a more controlled environment for the cells, aim to overcome these limitations [4].
References
[1] Agarose Wikipedia page: https://en.wikipedia.org/wiki/Agarose
[2] Protocol from Hyman Lab (MPI): Resources, Methods: https://hymanlab.mpi-cbg.de/hyman_lab/c-elegans/
[3] Kim E.et al. Long-term imaging of Caenorhabditis elegans using nanoparticle-mediated immobilization. PLoS One. (2013). https://www.ncbi.nlm.nih.gov/pubmed/23301069
[4] Ducret A. et al. A microscope automated fluidic system to study bacterial processes in real time. PLoS One. (2009). https://www.ncbi.nlm.nih.gov/pubmed/19789641
FAQ
Imaging non-adherent cells is difficult because they move. Living cells are motile, while fixed cells can move due to Brownian motion. It is necessary to prevent the movement of cells that are being imaged. This is especially needed when doing multichannel fluorescent microscopy, as the channels must be merged for colour overlays. Agar pads are used because they prevent both the motility of live cells and the Brownian motion of fixed cells. This makes them a suitable method for imaging multiple model organisms. These organisms include bacteria, yeast, worm embryos, adult worms, plants, and zebrafish embryos. They can also be combined with other methods, such as anaesthetics for C. elegans, to further improve immobilisation.
Agar pads are thin, flat layers of an agarose matrix. Although they are commonly referred to as "agar" pads, they are usually made from agarose. Agar is a polysaccharide substance obtained from red algae. Agarose is the main component of agar and is obtained after the agar has been purified. As a polymer, agarose creates a porous matrix. This structure traps cells, or molecules for molecular biology applications, preventing them from moving. Nutrient and gas exchange can still occur through this matrix, which makes the method compatible with live imaging. No toxicity for cells has been described as a result of growing in agar.
Yes, the percentage of agarose used to make the pads can be varied. Very standard percentages are from 0.5% to 2%. Higher percentages of agarose, up to 10%, have been used to immobilize adult worms. This method has the benefit of a lack of toxicity compared to other methods. Adult worms in these pads can be imaged for hours, provided the pad is properly sealed to avoid desiccation. It is also possible to replace agarose with other polysaccharides. For example, while 2% agar is standard for C. elegans embryo imaging, this can be replaced by 4% sucrose 0.1M NaCl for delicate early embryos or those sensitive to osmotic stress.
Agarose pads must be prepared so they are thin and flat. This allows them to be placed in a sandwich between a glass slide and a coverslip. The required thickness of the pad depends on the organism that will be imaged. One protocol shows how to prepare a 250µM thick pad using simple laboratory materials like glass slides and tape. The thickness of the tape is an important detail in this method. If the tape is too thin, it will result in very thin agar pads. These pads are often fragile and difficult to manipulate. This mounting protocol is also compatible with thicker pads, which may be needed for thicker specimens such as zebrafish embryos.
There are several main limitations when using agar pads for live cell imaging. One limitation is the impossibility of changing the media, which prevents actions like applying drugs during the experiment. Another risk is the pad drying out, also known as desiccation. For imaging experiments that take a long time, the limiting nutrient availability within the pad can also be a problem. These issues must be considered when planning experiments. Microfluidic technologies are intended to address these specific limitations. These newer technologies aim to ensure a more controlled environment for the cells being studied.